RCK is a software to infer protein-RNA preferences from RNAcompete experimental data.
RCK was developed by Yaron Orenstein
in Bonnie Berger's group
at Massachusetts Institute of Technology: MIT.
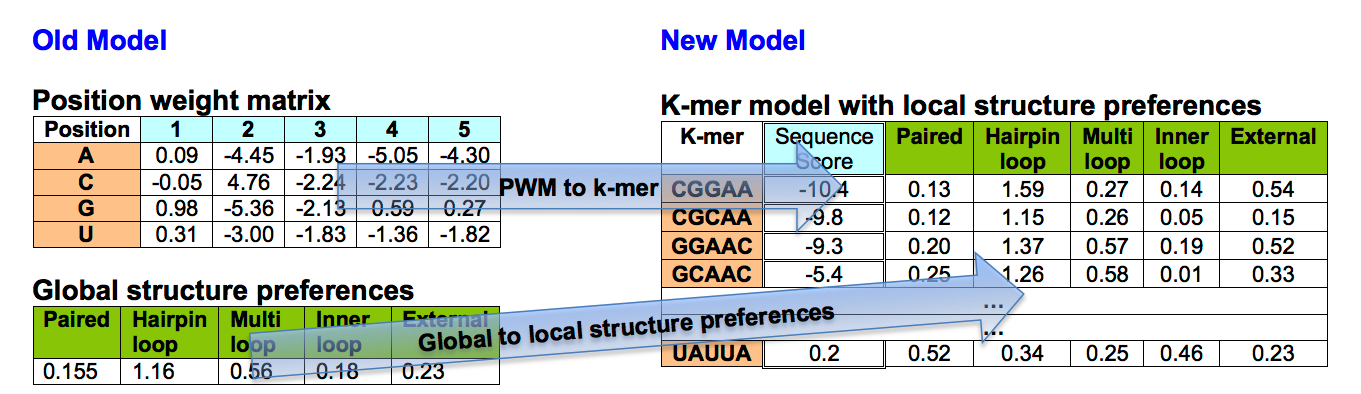
It extends RNAcontext by a k-mer model, in both sequence and structure.
Input and Output
For input and output description, see the original RNAcontext website.
The model output files have changed as the model has changed.
New features and parameters
Get the software
The package available here:
By default they are based on PHIME structure annotation: paired, hairpin, inner, multi and external.
PU models are based on paired and unpaired structure annotation.
RNAplfold implemented to produce probabilities for 4 structure contexts can be found here.
For training: see the original RNAcontext website.
And also: ./bin/rnacontext -w <min_width-max_width> -a <alphabet> -e <structure alphabet> -s <seed initialization>
-c <training sequences> -h <training probability vectors> -d <sequences to predict> -n <corresponding probability vectors>
-m <output dir> -b <max L-BFGS iteration>
Example run for training:
./bin/rnacontext -b 200 -w 4-5 -a ACGU -e PLMU -s 3 -c VTS1_training_sequences.txt -h VTS1_training_annotations.txt -d VTS1_test_sequences.txt -n VTS1_test_annotations.txt -o VTS1_demo -m ./outputs/
For prediction:
./bin/rnacontext -w <width-width> -a <alphabet> -e <structure alphabet> -d <sequences to predict> -n <corresponding probability vectors>
-l <model prefix> -m <dir of model and output> -b <max L-BFGS iteration>
Saves prediction results under <dir>/pred_<prefix>_<width>.txt
Example run for prediction:
./bin/rnacontext -w 5-5 -a ACGU -e PLMU -d VTS1_test_sequences.txt -n VTS1_test_annotations.txt -l VTS1_demo -m ./outputs/
Both modes output a PWM for visualization, and structure preference of top k-mer to pwm_* files in the output dir.