 Coev2Net
Coev2Net
Coev2Net is a general framework to predict, assess and boost confidence in individual interactions inferred from a HTP experiment. For every pair of interaction in the HTP screen, Coev2Net provides a score to assess their likelihood of being co-evolved from interacting homologous sequences. To do this, Coev2Net first predicts a likely interface model for the two proteins, by threading the sequences onto the best-fit template complex in our library. It then computes the likelihood of co-evolution of the two proteins (i.e. of the predicted interface) with respect to a probabilistic graphical model induced by the aligned interfaces of artificial homologous sequences. The graphical model is computed by solving a max-weight spanning tree problem over a graph of interface resiues, with the weights indicating the strength of correlation between the residues.
Supplementary Information for our paper "A computational framework to boost confidence in high-throughput protein-protein interaction datasets" is here.
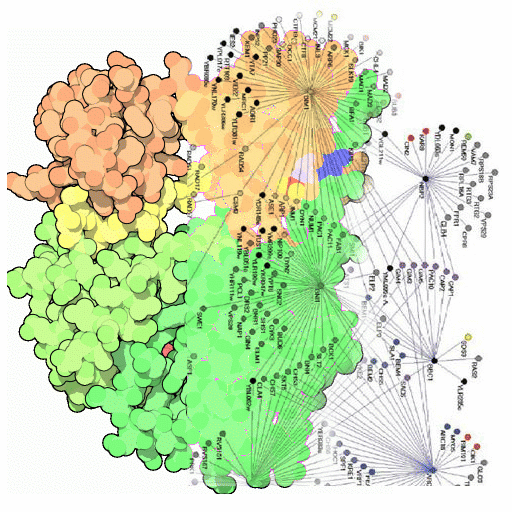
Coev2Net predictions for the MAPK network here.
Note that these sets do NOT contain PPI data used in training and testing the classifier (i.e., high-confidence interactions from Bandyopadhyay dataset).